Multi-bin Poisson#
[1]:
import json
import math
from pathlib import Path
import matplotlib.pyplot as plt
import numpy as np
import scrapbook as sb
from scipy.interpolate import griddata
import pyhf
from pyhf.contrib.viz import brazil
from pyhf.simplemodels import uncorrelated_background
[2]:
def plot_histo(ax, binning, data):
bin_width = (binning[2] - binning[1]) / binning[0]
bin_leftedges = np.linspace(binning[1], binning[2], binning[0] + 1)[:-1]
bin_centers = [le + bin_width / 2.0 for le in bin_leftedges]
ax.bar(bin_centers, data, 1, alpha=0.5)
def plot_data(ax, binning, data):
errors = [math.sqrt(d) for d in data]
bin_width = (binning[2] - binning[1]) / binning[0]
bin_leftedges = np.linspace(binning[1], binning[2], binning[0] + 1)[:-1]
bin_centers = [le + bin_width / 2.0 for le in bin_leftedges]
ax.bar(
bin_centers,
data,
0,
yerr=errors,
linewidth=0,
error_kw={"ecolor": "k", "elinewidth": 1},
)
ax.scatter(bin_centers, data, c="k")
[3]:
validation_datadir = "../../validation/data"
[4]:
with (
Path(validation_datadir).joinpath("1bin_example1.json").open(encoding="utf-8") as fp
):
source = json.load(fp)
model = uncorrelated_background(
source["bindata"]["sig"], source["bindata"]["bkg"], source["bindata"]["bkgerr"]
)
data = source["bindata"]["data"] + model.config.auxdata
init_pars = model.config.suggested_init()
par_bounds = model.config.suggested_bounds()
(
obs_limit,
exp_limits,
(poi_tests, tests),
) = pyhf.infer.intervals.upper_limits.upper_limit(
data, model, np.linspace(0, 5, 61), level=0.05, return_results=True
)
/srv/conda/envs/notebook/lib/python3.7/site-packages/pyhf/infer/calculators.py:352: RuntimeWarning: invalid value encountered in double_scalars
teststat = (qmu - qmu_A) / (2 * self.sqrtqmuA_v)
[5]:
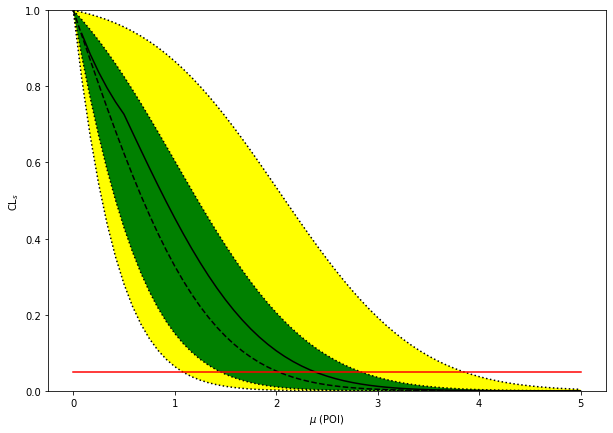
fig, ax = plt.subplots(figsize=(10, 7))
artists = brazil.plot_results(poi_tests, tests, test_size=0.05, ax=ax)
print(f"expected upper limits: {exp_limits}")
print(f"observed upper limit : {obs_limit}")
expected upper limits: [array(1.07644221), array(1.44922838), array(2.01932904), array(2.83213651), array(3.84750318)]
observed upper limit : 2.381026330918668
[6]:
source = {
"binning": [2, -0.5, 1.5],
"bindata": {
"data": [120.0, 145.0],
"bkg": [100.0, 150.0],
"bkgerr": [15.0, 20.0],
"sig": [30.0, 45.0],
},
}
my_observed_counts = source["bindata"]["data"]
model = uncorrelated_background(
source["bindata"]["sig"], source["bindata"]["bkg"], source["bindata"]["bkgerr"]
)
data = my_observed_counts + model.config.auxdata
binning = source["binning"]
nompars = model.config.suggested_init()
bonly_pars = list(nompars)
bonly_pars[model.config.poi_index] = 0.0
nom_bonly = model.expected_data(bonly_pars, include_auxdata=False)
nom_sb = model.expected_data(nompars, include_auxdata=False)
init_pars = model.config.suggested_init()
par_bounds = model.config.suggested_bounds()
print(init_pars)
bestfit_pars = pyhf.infer.mle.fit(data, model, init_pars, par_bounds)
bestfit_cts = model.expected_data(bestfit_pars, include_auxdata=False)
[1.0, 1.0, 1.0]
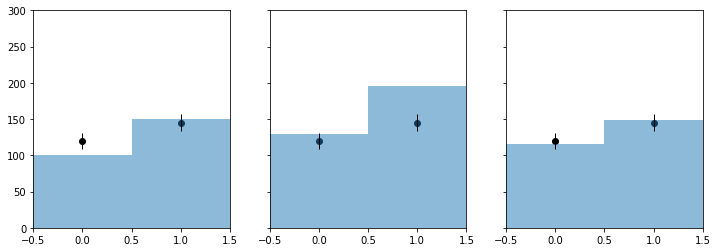
[7]:
f, axarr = plt.subplots(1, 3, sharey=True)
f.set_size_inches(12, 4)
plot_histo(axarr[0], binning, nom_bonly)
plot_data(axarr[0], binning, my_observed_counts)
axarr[0].set_xlim(binning[1:])
plot_histo(axarr[1], binning, nom_sb)
plot_data(axarr[1], binning, my_observed_counts)
axarr[1].set_xlim(binning[1:])
plot_histo(axarr[2], binning, bestfit_cts)
plot_data(axarr[2], binning, my_observed_counts)
axarr[2].set_xlim(binning[1:])
plt.ylim(0, 300);
[8]:
## DUMMY 2D thing
def signal(m1, m2):
massscale = 150.0
minmass = 100.0
countscale = 2000
effective_mass = np.sqrt(m1**2 + m2**2)
return [countscale * np.exp(-(effective_mass - minmass) / massscale), 0]
def CLs(m1, m2):
signal_counts = signal(m1, m2)
pdf = uncorrelated_background(
signal_counts, source["bindata"]["bkg"], source["bindata"]["bkgerr"]
)
try:
cls_obs, cls_exp_set = pyhf.infer.hypotest(
1.0, data, pdf, init_pars, par_bounds, return_expected_set=True
)
return cls_obs, cls_exp_set, True
except AssertionError:
print(f"fit failed for mass points ({m1}, {m2})")
return None, None, False
[9]:
nx, ny = 15, 15
grid = grid_x, grid_y = np.mgrid[
100 : 1000 : complex(0, nx), 100 : 1000 : complex(0, ny)
]
X = grid.T.reshape(nx * ny, 2)
results = [CLs(m1, m2) for m1, m2 in X]
[10]:
X = np.array([x for x, (_, _, success) in zip(X, results) if success])
yobs = np.array([obs for obs, exp, success in results if success]).flatten()
yexp = [
np.array([exp[i] for obs, exp, success in results if success]).flatten()
for i in range(5)
]
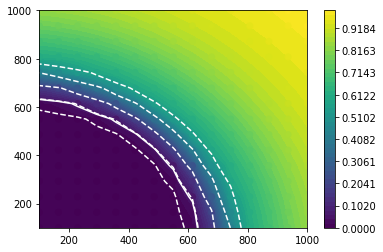
[11]:
int_obs = griddata(X, yobs, (grid_x, grid_y), method="linear")
int_exp = [griddata(X, yexp[i], (grid_x, grid_y), method="linear") for i in range(5)]
plt.contourf(grid_x, grid_y, int_obs, levels=np.linspace(0, 1))
plt.colorbar()
plt.contour(grid_x, grid_y, int_obs, levels=[0.05], colors="w")
for level in int_exp:
plt.contour(grid_x, grid_y, level, levels=[0.05], colors="w", linestyles="dashed")
plt.scatter(X[:, 0], X[:, 1], c=yobs, vmin=0, vmax=1);
[12]:
sb.glue("number_2d_successpoints", len(X))
Data type cannot be displayed: application/scrapbook.scrap.json+json